2024
Paper published on the optimality of the BMP pathway
A collaborative team from the University of Notre Dame and Purdue, and led by the Reeves lab at Texas A&M, has recently published a paper analyzing the optimality of the Bone Morphogenetic Protein (BMP) pathway, which is responsible for multiple cellular outcomes, including differentiation and cell death. In the paper, with first author Razeen Shaikh and titled “Optimal Performance Objectives in the Highly Conserved Bone Morphogenetic Protein Signaling Pathway,” we analyze how a model of the pathway can achieve specific desired behaviors, such as a fast response time and noise filtering. We found these behaviors, called “Performance Objectives” (POs), compete with each other, so that it is crucial for cells to balance these POs in the best way possible for their situation. We suggest that cells can alter the balance of these POs by regulating the concentrations of key proteins in the pathway. The paper has been published in the journal npj: systems biology and applications and can be found online here.
Paper submitted on the dynamics of transcription factors
Graduate student Sadia Dima is first author on a recently-submitted manuscript studying the dynamics of two crucial transcription factors in the early Drosophila embryo: Zelda and GAF. Sadia used a method called raster image correlation spectroscopy to measure the absolute concentration, binding, and mobility of these two transcription factors over a period of two hours in live embryos. With this work, she also was able to map out dose/response relationships for both of these transcription factors. The manuscript can be found online at here at the preprint server, bioRχiv.
Paper published on the dynamics of BMP signaling
A team of researchers from the Reeves lab, in collaboration with the Williams lab at NCSU, has recently published a paper analyzing the dynamics of the BMP pathway in early Drosophila embryos. The manuscript, co-first authored by former student Hadel Al Asafen and former postdoc Aydin Beseli, is titled “Dynamics of BMP signaling and stable gene expression in the early Drosophila embryo.” The is published in the Open Access journal Biology Open here. Update: see more in the press release here.
Reeves lab receives NSF funding
Dr. Reeves has led a team of researchers from Notre Dame and Purdue in securing funding from the NSF to study the Bone Morphogenetic Protein (BMP) signaling pathway across multiple species. The BMP pathway is highly conserved across the animal kingdom and is responsible for many different biological processes. In this project, we are asking the question of how the same signaling pathway can be responsible for multiple different kinds of outcomes. What is different between fruit fly embryos and human stem cells that the same pathway acts differently? A major goal of this work is to create a predictive mathematical model that is accurate to describe the behavior of the BMP pathway in three different species: fruit flies, zebrafish, and human cells. Read more here.
Paper on imaging of Cactus submitted
The Reeves Lab has recently submitted a paper, together with the Williams lab at NCSU, analyzing the distribution of the Cactus/IκBα, which is an inhibitor of the Dorsal/NF-κB pathway in Drosophila. The manuscript, titled “Spatiotemporal dynamics of NF-κB/Dorsal inhibitor IκBα/Cactus in Drosophila blastoderm embryos,” is first-authored by Dr. Allison Schloop (a recent graduate of the Reeves lab with her Ph.D. in Genetics). In this manuscript, we use a novel imaging technique in which the protein of interest, Cactus, is fused with a small protein that dynamically binds GFP. In this way, the distribution of Cactus can be inferred from fluorescent images in live embryos. We also used the advanced imaging techniques “Fluorescence Recovery after Photobleaching” (FRAP) and “Raster Image Correlation Spectroscopy” (RICS) to measure the presence of Cactus in the nucleus. The manuscript can be found online at here at the preprint server, bioRχiv.
Image analysis pipeline paper submitted
We have recently submitted a paper in collaboration with the Williams lab at NCSU demonstrating a new pipeline for analyzing images of the microscopy technique “Fluorescence Recovery after Photobleaching” (FRAP). In this technique, an area of fluorescence is bleached (goes dark) using strong laser power, and the dynamics of recovery of that darkened area can tell you how mobile the fluorescent molecules are. In this manuscript, titled “FRAP analysis: Measuring biophysical kinetic parameters using image analysis” and authored by Ph.D. student Sharva Hiremath (co-advised by Dr. Williams and Dr. Reeves), we describe the formulation and use of image analysis algorithms to extract quantitative data from these FRAP images. The manuscript can be found online at here at the preprint server, arχiv.
Paper on the mobility of Dorsal submitted
A team of researchers from the Reeves lab, in collaboration with the Sozzani lab at NCSU, has recently (re)submitted a paper in which we measure the mobility of Dorsal in early Drosophila embryos. The manuscript, co-first authored by former student Hadel Al Asafen, Dr. Sozzani’s former graduate student Natalie Clark, and former postdoc Etika Goyal, is titled “Dorsal/NF-κB exhibits a dorsal-to-ventral mobility gradient in the Drosophila embryo.” In this work, we use the advanced imaging technique known as “Raster Image Correlation Spectroscopy” (RICS) to measure the mobility and binding of Dorsal, and we find that Cactus is likely entering the nucleus to prevent Dorsal/DNA interactions. The manuscript can be found online at here at the preprint server, bioRχiv.
2023
Allison Schloop has graduated with her Ph.D.!
Today Allison Schloop graduated from NCSU with a PhD in Genetics. Her thesis is titled “Dynamics of Inhibitory Regulation of the Drosophila NF-κB/Dorsal Gradient.” Congratulations Allison!
Reeves lab secures NSF funding
Dr. Reeves has received funding from the NSF to study Drosophila female germline stem cell (GSC) differentiation. The GSC receives Bone Morphogenetic Protein (BMP) pathway signals from nearby cells, which tell the GSC to stay a stem cell. After cell division, one of the daughter cells receives less BMP signal, and so it differentiates into the eventual egg cell. This pathway is highly regulated to make sure nothing goes wrong in this process. Our work will study the regulation from a quantitative standpoint so that, eventually, we will know how to control the differentiation process. Read more here.
Ph.D student Razeen Shaikh earns departmental fellowship!
Razeen Shaikh was recently awarded the Paul & Ellen Deisler Chemical Engineering Fellowship for 2023-2024. As described by the department, “This award is a testament to Razeen’s excellent work and success while at Texas A&M.” Congratulations Razeen!
Systems biology optimzation paper accepted!
A recent manuscript submission from the Reeves lab has been accepted to the journal Bioinformatics! The paper, co-first authored by recently-graduated Dr. Prasad Bandodkar and current Ph.D. student Razeen Shaikh, is titled “ISRES+: An improved evolutionary strategy for function minimization to estimate the free parameters of systems biology models.” The published paper can be found here. Read more in this press release.
2022
Prasad Bandodkar has graduated with his Ph.D.!
Today Prasad Bandodkar defended his PhD thesis, titled “Computational studies of the patterning systems in early Drosophila embryos.” After graduation, he will be working as an Image Analysis Scientist with Bio-Techne in the San Francisco Bay Area. Congratulations Prasad!
Paper published on quantificaiton of protein binders
In a recent collaboration with the Rao Lab at NCSU, the Reeves lab has recently published a paper in ACS Omega titled, “Experimental and analytical framework for ‘mix-and-read’ assays based on split luciferase.” Former postgraduate worker Nikki McArthur, co-advised by Dr. Reeves and Dr. Rao, was the first author. You can read more about the paper, and how it integrates into the research being done by the Reeves lab, in this press release.
Razeen Shaikh wins best poster!
Ph.D. student Razeen Shaikh won best poster for her poster presentation in the 9th annual Chemical Engineering Graduate Student Association (ChEGSA) Symposium. The symposium showcases the research from the Chemical Engineering department here at Texas A&M. Congratulations Razeen!
Shelby Morton joins the lab!
Shelby Morton, a first-year Ph.D. student from the Interdisciplinary Program in Genetics, has joined the lab. He will be working on mesasuring the dynamics of the BMP gradient in live embryos. He will be focusing on in particular on the mechanisms that connect transient signaling to sustained gene expression. Welcome Shelby!
2021
Sadia Dima joins the lab!
Earlier this month, first-year Ph.D. student Sadia Siddika Dima joined the lab. She will be working on global measurements of the Dorsal gradient in live embryos. She will be focusing on measuring the Dorsal gradient in mutant embryos in particular. Welcome Sadia!
GSA spotlight on Ph.D. student Elle Rooney!
Last month, an interview with Ph.D. student Lossie (Elle) Rooney was posted to “Genes to Genomes,” a blog maintained by the Genetics Society of America (GSA). This blog post is part of a series in which they spotlight members of GSA’s Early Career Leadership Committees. Read the blog here. Great work Elle!
Two students join the lab!
Last week, first-year Chemical Engineering Ph.D. students Hung-Yuan (Zeke) Chen and Razeen Shaikh joined the lab. They will be working on experimental and modeling approaches to studying the BMP pathway. Welcome Zeke and Razeen!
2020
Collaboration project awarded T3 funding
A recently-established collaboration with Dr. James Erickson and Dr. Aref Zarin (both in the Biology department) has been selected for funding through the T3 mechanism! The collaborative project, titled “Transvection As A General Means To Synchronize Gene Expression,” focuses on regulation of a gene called Sex lethal (Sxl), which determines whether a developing Drosophila embryo becomes male or female. As a means of regulation of Sxl (or any other gene), transvection is when DNA from one chromosome activates DNA on the paired chromosome. This project will investigate novel aspects of transvection.
Collaboration project with NCSU investigators awarded RISF funding
Reeves Lab collaborators at NC State University — Julio Belmonte (Physics), Mary Elting (Physics), and Caroline Laplante (Molc. Cellular Biosci) — have been awarded funding through NC State’s “Research Innovation and Seed Funding” program. The collaborative project focuses on the dynamics of cellular membrane tension in the 3 hr old Drosophila embryo. The funding will provide continued support for our co-advised postdoc Sophia Webster. Congratulations Julio, Mary, and Caroline!
Paper published on embryo penetrating peptides
The Rao and Menegatti labs (CBE, NCSU), in a collaboration between the Reeves lab, have published a paper in Bioconjugate Chemistry. In this work, Dr. John Bowen screened for membrane penetrating peptides using Drosophila embryos. After selecting the best-performing peptide, he showed that it can penetrate not only Drosophila embryos, but also human stem cells. Dr. John Bowen is a recent Ph.D. graduate from the CBE department at NCSU, co-advised by Drs. Rao and Menegatti. The Reeves lab lent expertise in Drosophila biology. Allison Schloop from the Reeves lab was co-author. Congratulations John and Allison! Read more here.
Reeves lab moves to Texas A&M University!
Even during the pandemic of 2020, there can be some good news! We are pleased to announce that the Reeves Lab has moved to Texas A&M in College Station, TX. We have joined the Chemical Engineering Department, and are also part of the Interdisciplinary Graduate Program in Genetics. Texas A&M is a fantastic university with the largest student population in the country. We are excited about the opportunities that lie ahead.
Self-cleaving ribozymes paper published
One of the papers described below (see news post from Aug 19, 2019) has been accepted for publication at PLoS One. In this paper, Dr. Thomas Jacobsen, with Gloria Yi, Hadeel Al Asafen, and Dr. Ashley Jermusyk, created customizable RNA sequences that would degrade themselves. While this may sound counter-productive, these sequences could be tuned to self-degrade at given rates, which will allow researchers to tune the level of protein produced by the RNA. Dr. Jacobsen received his Ph.D. in May 2019 and was co-advised between Dr. Reeves and Dr. Chase Beisel (Helmholtz Institute). Read more here.
Paper published on Robustness of the Dorsal gradient
One of the papers described below (see news post from Aug 19, 2019) has been accepted for publication at PLoS Computational Biology. In this paper, co-first authors Hadeel Al Asafen and Prasad Bandodkar investigate how the Dorsal gradient system is affected by changing the amount of Dorsal given to the embryo by the mother. Hadeel performed experiments in live Drosophila embryos to measure how much the Dorsal gradient changes in different embryos. Prasad performed computational analysis, and importantly, found that three mechanisms that the Reeves lab has previously published on are responsible for the robustness. For the full story, see here. Dr. Sophia Carrell, Allison Schloop, and Jeramey Friedman were co-authors.
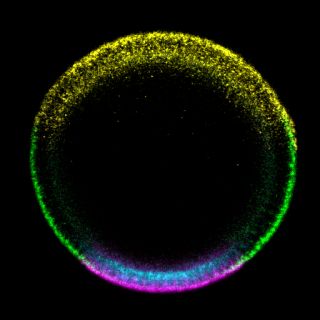
Our review of the Dorsal gradient is available online!
The review paper described below (see news post from Aug 26, 2019) is now available online. An image of one of our embryos is part of the cover art! See here.
Paper published in Developmental Biology
One of the papers described below (see news post from Aug 19, 2019) has been accepted for publication at Developmental Biology. In this paper, we review the implications of the shuttling mechanism that we previously published. We also suggest that the shuttling mechanism can be used predictively, and present new data in which the shuttling mechanism may be acting in cross-talk between the Dorsal and Dpp systems. Read more here.
Paper published in Developmental Dynamics
Ph.D. student Prasad Bandodkar has analyzed the action of a coherent feedforward loop activated by a morphogen gradient. He has found, for both a generic morphogen and the Dorsal morphogen, that the structure of the coherent feedforward loop naturally acts to stabilize variations in the input (morphogen). Feedforward loops are very commonly studied, but they have almost exclusively been studied either varying in time (but not space), or varying in space (but at steady state). However, most morphogens are not at steady state. In particular, we have long known the Dorsal gradient is dynamic in time. Co-factor Twist, as the intermediate node in the feedforward loop, together with the time-varying levels of the pioneering factor Zelda, stabilize the Dorsal gradient’s dynamics. Read more here.
2019
Reeves lab submits major review article
The Reeves lab has focused on quantitative and computational aspects of the Drosophila Dorsal/NF-κB network, and has published or submitted several papers on this topic since 2015. Furthermore, as a postdoc, Dr. Reeves had also published highly-cited articles on quantitative discoveries of the shape and dynamics of the Dorsal gradient. Therefore, we are pleased to have just submitted a comprehensive review article on this system, with authors Allison Schloop and Prasad Bandodkar. The article has a specific focus on the quantitative and computational aspects of the Dorsal gradient network. This work should go to press to appear in March, 2020.
Reeves lab posts three manuscripts to bioRχiv
The Reeves lab has posted three manuscripts to bioRχiv, the most prominent preprint server for biological research. In the first manuscript, in collaboration with Dr. Chase Beisel’s lab and with lead author Dr. Thomas Jacobsen, we characterize the use of self-cleaving ribozymes for regulating gene expression. The work was performed in mammalian cells and Drosophila embryos, demonstrating the versatility of this gene regulation tool in multiple organisms. Read more here.
In the second manuscript, we describe in detail the mechanism and phenotypes of facilitated diffusion (or shuttling) of Dorsal by Cactus in the early Drosophila embryo. This is a diffusion mechanism we have published previously, showing that, counterintuitively, it causes Dorsal to accumulate on one side of the embryo. Furthermore, we suspect it is necessary to achieve the peak levels of Dorsal signaling in the embryo. In the new manuscript, we also show that it is important for cross-talk with the BMP pathway. Read more here.
In the third study, we show that the Dorsal gradient is robust to changes in the initial amount of Dorsal in the embryo. Furthermore, we show that both shuttling (referred to above) and the presence of the inhibitor Cactus in the nucleus (see also our previous work here), are required for this robustness. Read more here.
Dr. Reeves publishes in JBE
Dr. Reeves had a paper published in the “Emerging leaders in biological engineering” thematic series of the Journal of Biological Engineering. Gene regulation is paramount in all aspects of biology, and one of the main motifs in gene regulation is the feedforward loop. Inspired by engineering principles found in man-made systems, Dr. Reeves shows that feedforward control in biology can be aided by adding feedback control to the gene regulation. He also shows that genetic feedback loops are present in E. coli, which was previously unreported. Read more here.
Reeves lab secures NSF funding
Dr. Reeves has received funding from the NSF to begin quantitative and computational work on the Dorsal/NF-κB signaling network in the Drosophila embryo. One of the paradoxes of “systems biology” is that mathematical models are needed to help interpret data on such a large scale, yet the models themselves are so large that many their unknowns hinder interpretation. The project will break down the individual processes in the system to focus on them individually. Once those are well-understood, the full system can be elucidated. Read more here.
Thomas Jacobsen earns his Ph.D!
Tom Jacobsen, co-advised by Dr. Reeves and Dr. Chase Beisel (Helmholtz Inst.) successfully defended his Ph.D. work today. His projects focused on methods of gene regulation using CRISPR and self-cleaving ribozymes. He has been an invaluable member of the Reeves lab in both his mentoring of younger graduate students and undergraduates, as well as providing research expertise to aid in compile multiple manuscripts as contributing author. Congratulations Tom!
2018
Hadeel Al Asafen wins image contest!
Graduate student Hadeel Al Asafen has won first place in the NCSU 2018 Envisioning Research contest in the student/postdoc category. Her video can be seen on youtube here. It depicts the dynamics Dorsal gradient when the nuclei are dividing multiple times. Read more about the contest here.
Allison Schloop joins the lab!
We are pleased to announce that a new student from the Genetics Program, Allison Schloop, has joined the lab. Allison earned a bachelor’s degree in Biochemistry (and a minor in math) from St. Lawrence University. She will be working on the role of feedforward and feedback loops in the Dorsal gradient system.
Reeveslab has posted two new manuscripts on bioRχiv
Two recent studies performed in the lab have now been uploaded to the biology preprint server, bioRχiv. In the first study, we have used a panel of wild-caught, naturally-varying fly lines to infer relationships between proteins that act very early in embryo development. Read more here.
In the second study, we have used advanced, quantitative imaging methods to measure how proteins move in the early Drosophila embryo. Understanding protein movement is a major piece in understanding how tissues develop. Read more here.
Reeveslab welcomes four new members!
This semester, we are pleased to welcome Etika Goyal, Christopher McCallough, Prasad Bandodkar, and Lossie Rooney to the lab. Etika and Christopher are postdocs from Vellore Institute of Technology and University of Illinois at Chicago, respectively. Prasad is most recently from IIT, Delhi (Master’s degree), and Lossie recently became a PhD student after studying here at NCSU. See /people.php”>here for more on Reeveslab members.
2017
Reeveslab’s fly embryo image featured on BPoD
Biomedical Picture of the Day has featured an image of early fruit fly embryos from our recent work. The image contains two embryos: the one on the left is 1-1.5 hrs younger than the one on the right. During that time span, the transcription factor Dorsal diffuses through the embyro to accumulate on the ventral side. Read more here. Image credit: Sophie Carrell.
NSF highlights our recent work
Our NSF-funded research, which was recently published in Development, has been highlighted on the NSF website. See also our blog post regarding the work.
Our recent publication in the news
Our manuscript recently accepted in Development has been highlighted in a recent news release from NC State. The title of the news article is “Researchers Find Diffusion Plays Unusual Signaling Role in Drosophila Embryos,” and can be found here. (It has also been re-posted on various news outlets, such as here and here.)
ReevesLab is looking for one postdoc
Dr Reeves is looking for one postdoc to join the lab. The project will focus on studying the subtle changes in tissue patterning among a panel of naturally-varying fly lines. These lines have subtle genomic differences, which can be correlated to the changes in gene expression pattern. For more information, see the job posting here.
UPDATE: this project has been filled. Reeveslab is no longer looking for postdocs.
Manuscript accepted in Development!
Our study on the mechanism of Dorsal gradient formation has recently been accepted for publication at Development. The study was led by co-first authors Sophie Carrel and Mike O’Connell, and assisted by current student Tom Jacobsen, and two former undergraduate researchers Amy Pomeroy (Allen) and Stephanie Hayes (Smith).
ReevesLab is looking for two postdocs
Dr Reeves is looking for two postdocs to join the lab on two different projects. The first project will focus on the design of synthetic genetic network motifs in the early fruit fly embryo.
The second project will focus on studying the subtle changes in tissue patterning among a panel of naturally-varying fly lines. These lines have subtle genomic differences, which can be correlated to the changes in gene expression pattern. This job posting will open soon, so check back for details.
UPDATE: the first project has been filled, and the second project is open.
Dr Reeves receives NIH funding with Drs Rao and Williams
Dr Reeves has received funding from the NIH to begin work on two new projects. In the first, Dr Reeves and Dr Balaji Rao will collaborate to engineer novel sensors to detect RNA and short-lifetime proteins in living tissues. They will use the new technology to study the dynamics of regulatory loops in live tissues and cells.
In the second project, Dr Reeves and Dr Cranos Williams will collaborate to study regulation of signals and decision-making in developing tissues using natural variation. The lab of Dr Trudy Mackay will contrbute fly lines and expertise to the project.
Dr Reeves and Dr Hrischuk publish a second review paper
In the paper, Drs Reeves and Hrischuk review published “standard engineering principles” and find that the cell embodies each of these principles (save one). Additionally, they publish a novel list of Chemical Process Control Engineering Principles, and find the cell embodies these as well. The implication is there could be benefits from studying biology from an engineering point of view.
2016
Sophie Carrell earns PhD
Today Sophie has defended her doctoral dissertation! Sophie has published a paper on the microscopy of embryo cross sections, and has another paper on the formation of the Dorsal gradient under revision (see here). She also initiated other projects, including how the relative dosages of Dorsal and Cactus affect patterning and how the Dorsal pathway interacts with the Dpp pathway. Congratulations Dr Carrell!
Ashley Jermusyk publishes paper on synthetic network
Former PhD student and recent graduate Ashley Jermusyk (now a postdoc at the NCI) just had her paper accepted for publication in BMC Systems Biology. She created, analyzed, and refined a synthetic negative feedback loop in the early Drosophila melanogaster embryo. Read more here. Congratulations Ashley!
Ashley Jermusyk earns PhD
Ashley has successfully defended her PhD work! In her time in my lab, she worked on several different projects, including a synthetic negative feedback loop, natural variation of gene expression patterns, and ribozyme-mediated control of gene expression. Congratulations Dr Jermusyk!
Sophie Carrell wins 3rd place at the Schoenborn Symposium
Sophie has won the third place prize at the Schoenborn Symposium for her talk on the formation of the Dorsal gradient in the early Drosophila embryo. The Schoenborn Symposium is an annual research symposium showcasing the body of graduate research in the Department of Chemical and Biomolecular Engineering here at NC State. Congratulations Sophie!
Dr. Reeves and Dr. Curtis Hrischuk publish review paper
Dr. Reeves has published a a review paper in the Computational Biology Journal on the relationship between the fields of systems engineering and systems biology. The crucial observation is that cells constitute embedded computing systems. Check it out!
2015
Ashley Jermusyk publishes chapter on transcription factor networks
Ashley has published a review chapter in the Encyclopedia of Cell Biology. Congratulations Ashley!
Thomas Jacobsen awarded GAANN Fellowship
Tom, a graduate student co-advised by Dr. Beisel and Dr. Reeves, has been awarded a fellowship sponsored by the Graduate Assistance in Areas of National Need (GAANN) program in the area of Biotechnology. Congratulations Tom!
Michael O’Connell publishes paper on the Dorsal gradient
Mike has published a modeling paper in the journal PLoS Computational Biology. His work potentially reveals new roles for Dorsal’s inhibitor, Cactus. Congratulations Mike!
2014
Sophie Carrell publishes paper on embryo imaging
Sophie has published a methods protocol in the journal Methods in Molecular Biology. She has refined a method for manual dissection of fixed Drosophila embryos, as well as a method for upright imaging of live embryos. Congratulations Sophie!
Reeves and Sozzani awarded RISF funding
Dr. Reeves and Dr. Rosangela Sozzani (Plant Biology) have been awarded funding through NC State’s “Research and Innovation Seed Funding” program. During the next year, they will work on acquiring data on distribution and mobility of signaling molecules in the early Drosophila embryo.
Dr. Reeves and Dr. Beisel awarded NSF funding
Dr. Reeves and colleague Dr. Chase Beisel (also in CBE) receive NSF funding to build and tune synthetic circuits in early Drosophila embryos. Read more here.
2013
Michael O’Connell awarded GAANN Fellowship
Mike O’Connell, a graduate student in the Reeves lab, has been awarded a fellowship sponsored by the Graduate Assistance in Areas of National Need (GAANN) program in the area of Scientific Computing. Congratulations Mike!
Reeves and Deiters awarded RISF funding
Dr. Reeves and Dr. Alexander Deiters (Chemistry) have been awarded funding through NC State’s “Research and Innovation Seed Funding” program. During the next year, they will work on acquiring data on regulation of gene expression in the early Drosophila embryo.
Dr Reeves receives NSF CAREER Award
Dr Reeves has been selected to receive the Faculty Early Career Development (CAREER) Award from the National Science Foundation. From the NSF webpage: This is one of “the National Science Foundation’s most prestigious awards in support of junior faculty who exemplify the role of teacher-scholars through outstanding research, excellent education and the integration of education and research within the context of the mission of their organizations. Such activities should build a firm foundation for a lifetime of leadership in integrating education and research.” Read more here.
2012 and prior
Dr Reeves’ fly embryo image featured on BPoD
Biomedical Picture of the Day has featured an image of an early fruit fly embryo taken by Dr Reeves. This cross section of a 2.5 hr old embryo reveals the expression of five different genes in a spatial pattern. Read more here.
Dr Reeves selected as NCSU nominee for Pew Award
The Pew Scholars program grants 4-year awards to young professors in the biomedical sciences. Each school that participates in the competition can nominate one individual. In this round of competition, 22 Pew Scholars will be named.
Job posting for undergraduate research
I am currently looking for an undergraduate researcher to help set up my lab and continue working through the Spring 2011 and perhaps into the next year.
Start date for Dr Reeves changed to August 2010
I spoke with the department and together we decided to push my start date back to August of 2010. I will then be recruiting students and/or postdocs to begin that fall semester.
ABOVE: Cover art for the volume of Current Topics in Developmental Biology that our review of the Dorsal gradient was published in. BELOW: larger version of our image that was used in the cover art. Image credit: Silu Zhang