Click here for full list of publications.
Note: asterisk indicates shared first authorship.
Bandodkar et al., 2023
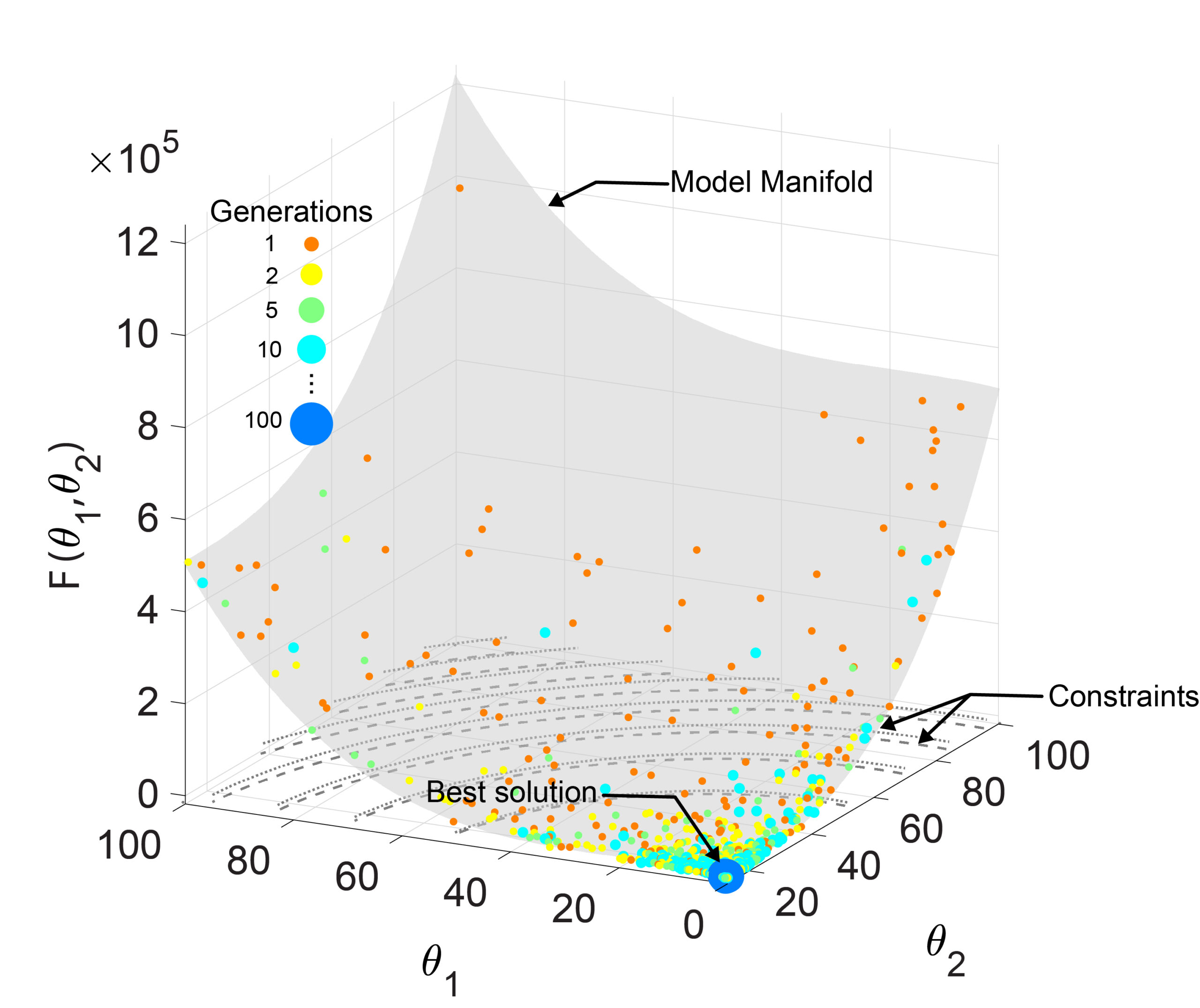
Methods to optimize models to data.
Bandodkar, P. U.*; Shaikh, R. R.* & Reeves, G. T. (2023). ISRES+: An improved evolutionary strategy for function minimization to estimate the free parameters of Systems Biology models. Bioinformatics, 39 (7):btad403. [Read more in this press release.]
McArthur et al., 2020
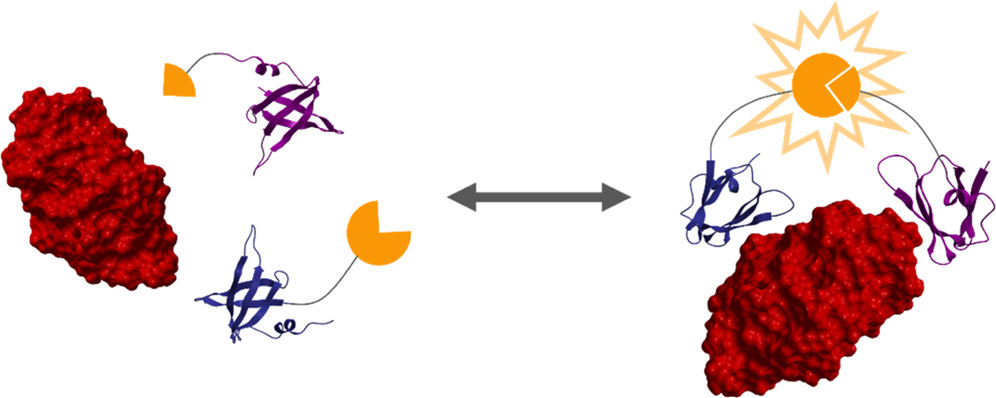
Simple, quantitative readouts of protein concentration.
McArthur, N.; Cruz-Teran, C.; Thatavarty, A.; Reeves, G. T. & Rao, B. M. (2022). Experimental and Analytical Framework for “Mix-and-Read” Assays Based on Split Luciferase. ACS Omega, 7: 24551-24560. doi: 10.1021/acsomega.2c02319. [Read more in this press release.]
Schloop et al., 2020
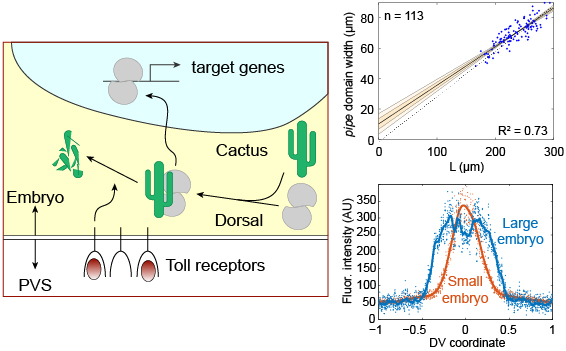
A comprehensive review of the Dorsal Gradient system.
Schloop, A. E.; Bandodkar, P. U. & Reeves, G. T. (2020). Formation, interpretation, and regulation of the Drosophila Dorsal/NF-κB gradient. Current Topics in Developmental Biology 137: 143-191.doi: 10.1016/bs.ctdb.2019.11.007 [Invited review.]
Jacobsen et al., 2020
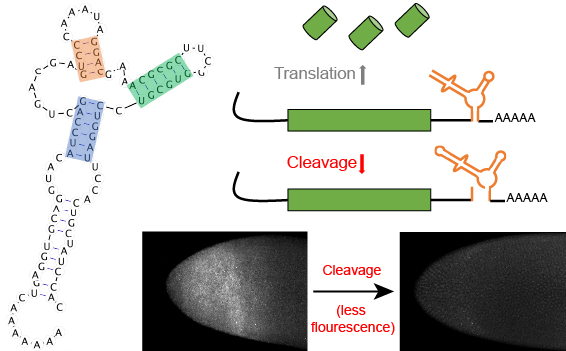
Self-degrading mRNA tools for tunable gene expression.
Jacobsen, T.; Yi, G.; Al Asafen, H.; Jermusyk, A. A.; Beisel, C. L. & Reeves, G. T. (2020). Tunable self-cleaving ribozymes for modulating gene expression in eukaryotic systems. PLoS One, 15: e0232046. doi: 10.1371/journal.pone.0232046
Bandodkar et al., 2020
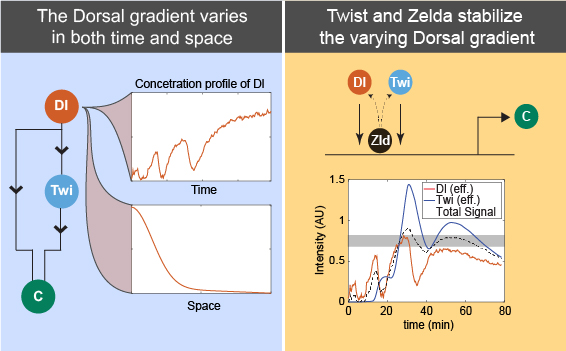
Feedforward control stabilizes morphogen gradients that vary in both space and time.
Bandodkar, P. U.; Al Asafen, H. & Reeves, G. T. (2020). Spatiotemporal control of gene expression boundaries using a feedforward loop. Dev Dyn, 249: 369-382. doi: 10.1002/dvdy.150 [Invited for a special issue: “50 Years of Positional information in Development, Disease, and Regeneration.”]
Al Asafen et al., 2020
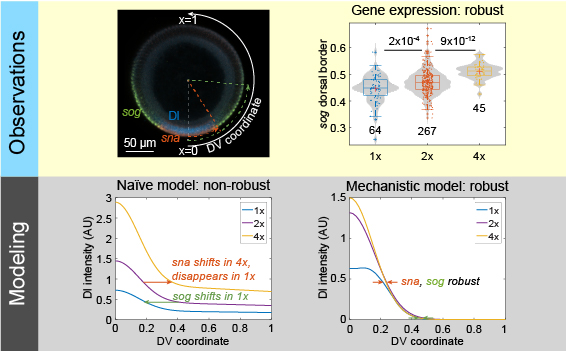
Dorsal target gene expression is unexpectedly robust. Mechanistic modeling reveals why.
Al Asafen, H.; Bandodkar, P. U.; Carrell-Noel, S.; Schloop, A. E.; Friedman, J. & Reeves, G. T. (2020). Robustness of the Dorsal morphogen gradient with respect to morphogen dosage. PLoS Computational Biology, 16: e1007750. doi: 10.1371/journal.pcbi.1007750
Reeves, 2019
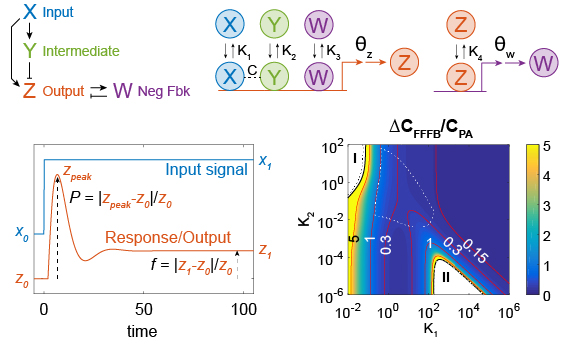
Combining feedforward and feedback control is beneficial in both man-made and biological systems.
Reeves, G. T. (2019). The engineering principles of combining a transcriptional incoherent feedforward loop with negative feedback. Journal of biological engineering, 13: 62. doi: 10.1186/s13036-019-0190-3 [Invited for a special issue: “Emerging leaders in Biological Engineering.”]
Carrell et al., 2017
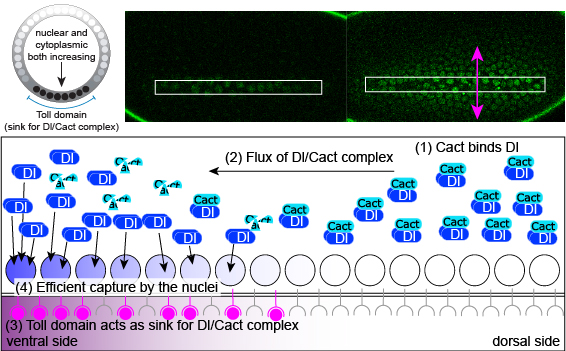
A signaling gradient goes against the grain: diffusion causes accumulation instead of spreading.
Carrell, S. N.*; O’Connell M. D.*; Jacobsen, T.; Pomeroy, A. E.; Hayes, S. M. & Reeves, G. T. (2017). A facilitated diffusion mechanism establishes the Drosophila Dorsal gradient. Development, 144: 4450-4461. doi: 10.1242/dev.155549
Jermusyk et al., 2016
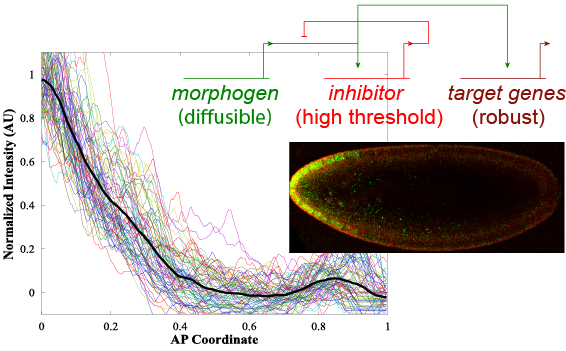
A synthetic negative feedback loop has unexpected consequences.
Jermusyk A. A.; Murphy, N. P. & Reeves, G. T. (2016). Analyzing negative feedback using a synthetic gene network expressed in the Drosophila melanogaster embryo. BMC Syst Biol, 10: 85. doi: 10.1186/s12918-016-0330-z
Jermusyk and Reeves, 2016
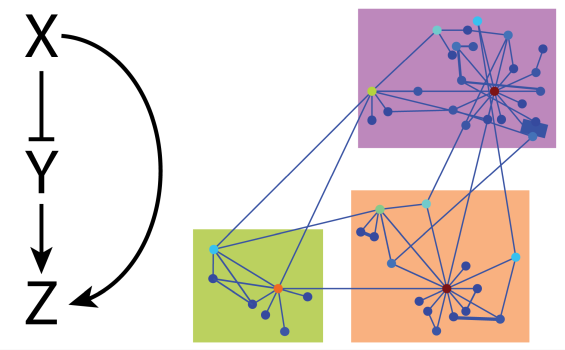
Transcription factor networks show similar global properties to man-made networks.
Jermusyk A. & Reeves, G. T. (2016). Transcription Factor Networks. Encyclopedia of Cell Biology, 4: 63-71. doi: 10.1016/B978-0-12-394447-4.40010-6
O’Connell and Reeves, 2015
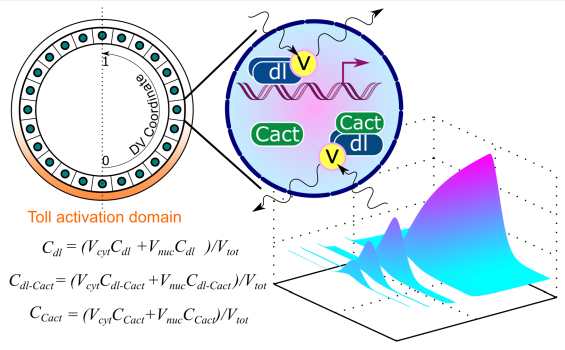
Computational modeling of the Dorsal gradient suggests that Dorsal/Cactus complex is present in the nucleus.
O’Connell, M. D. & Reeves, G. T. (2015). The presence of nuclear Cactus in the early Drosophila embryo may extend the dynamic range of the Dorsal gradient. PLoS Comput Biol, 11: e1004159. doi: 10.1371/journal.pcbi.1004159
Reeves et al., 2012
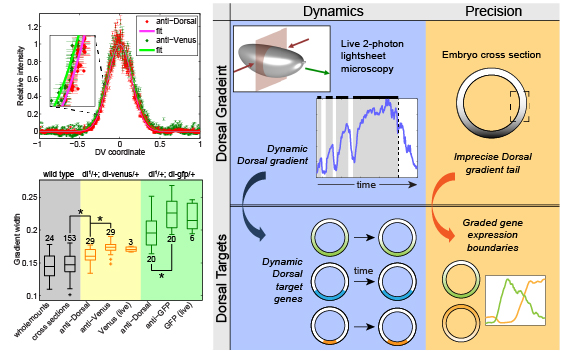
Live imaging of the Dorsal gradient found it to be highly dynamic.
Reeves, G. T.*; Trisnadi, N.*; Truong, T. V.; Nahmad, M.; Katz, S. & Stathopoulos, A. (2012). Dorsal-Ventral Gene Expression in the Drosophila Embryo Reflects the Dynamics and Precision of the Dorsal Nuclear Gradient. Dev Cell, 22: 544-557. doi: 10.1016/j.devcel.2011.12.007
Liberman et al., 2009
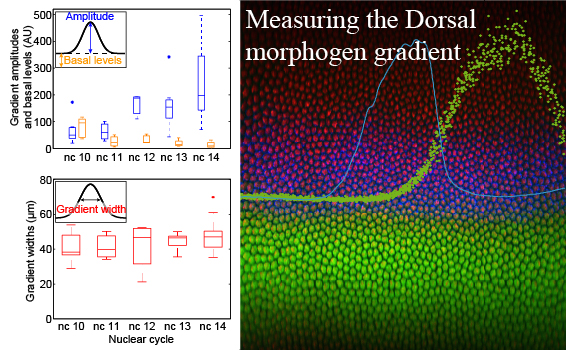
Detailed, quantitative measurements of the Dorsal gradient found it to be surprisingly narrow.
Liberman, L. M.*; Reeves, G.T.* & Stathopoulos, A. (2009). Quantitative imaging of the Dorsal nuclear gradient reveals limitations to threshold-dependent patterning in Drosophila. Proc Natl Acad Sci U S A, 106: 22317-22322. doi: 10.1073/pnas.0906227106.
Reeves and Stahopoulos, 2009
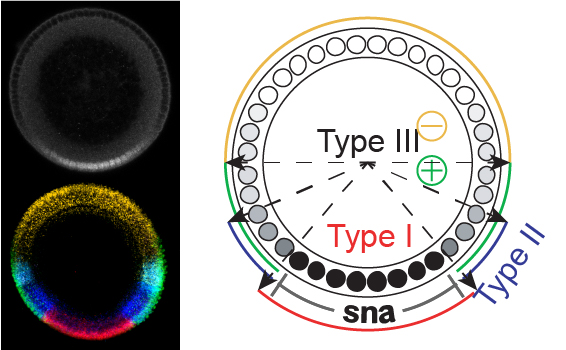
The Dorsal gradient divides the embryo into roughly three domains.
Reeves, G.T. & Stathopoulos, A. (2009). Cold Spring Harb Perspect Biol, “Perspectives on Generation and Interpretation of Morphogen Gradients.” doi: 10.1101/cshperspect.a000836.